 - A lightweight integration environment for model annotation
- A lightweight integration environment for model annotation
Developers: A.L. Lister, M. Pocock, A. Wipat
Official project page: Project: Saint
Download: Saint
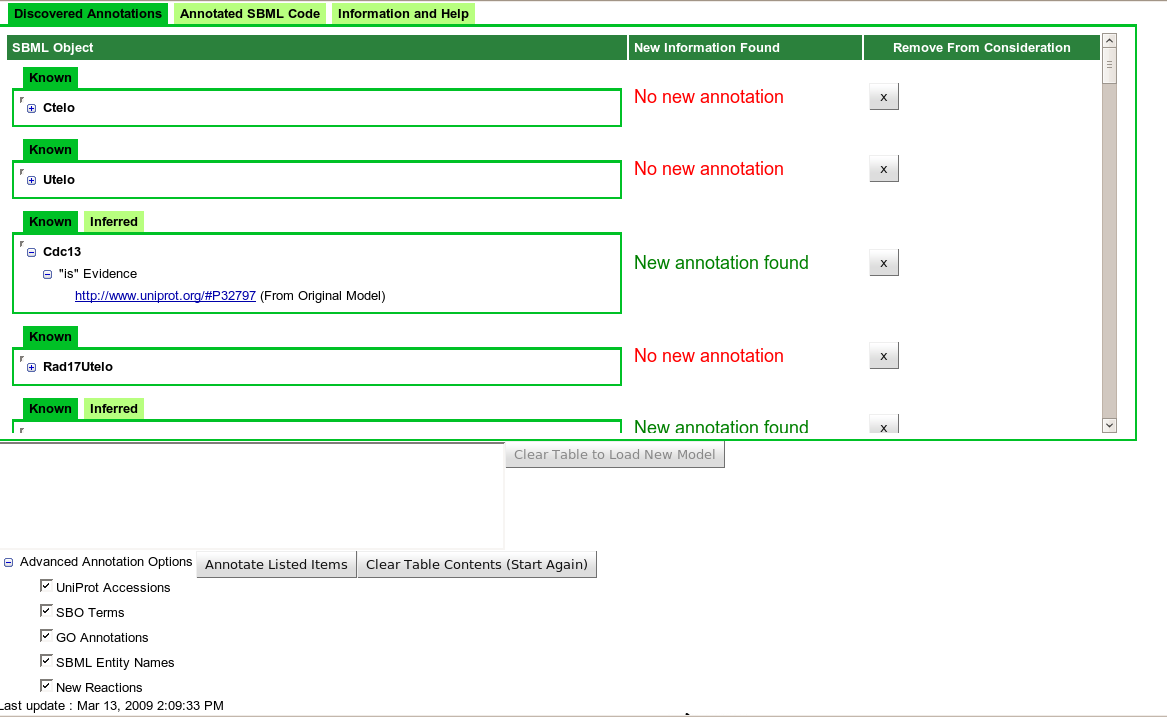 |
| Figure 1: "Discovered Annotations" tab after querying remote data sources. |
The creation of accurate quantitative biological models in languages such as SBML or CellML is a time-intensive manual process. Modellers need to know and understand both the systems they are modelling and the intricacies of the modelling language. However, the amount of relevant data for even a relatively small and well-scoped model is overwhelming. Saint, an automated annotation integration environment, aims to aid the modeller and reduce development time by providing extra information about any given model in an easy-to-use interface. Saint accepts SBML-formatted files and integrates information from multiple databases automatically. Any new information that the user agrees with is then automatically added to the model.
The initial functionality of Saint allows the annotation of already-extant species and suggests additional interactions. The user uploads their model, and the portions of the model recognized by Saint are then displayed using a tabular structure (see Figure 1). The user can then remove any items they are not interested in annotating. For instance, some terms such as "sink" are modelling artefacts and do not correspond to genes or proteins. Therefore, the user would normally wish to delete this from the search space to prevent any possible matches with actual biological species of a similar name. Once the user is satisfied with the list of items to be annotated, the model is submitted using the "Annotate Listed Items" button at the bottom of the table. A summary of the annotation returned by Saint is then added to the main table. The user can then remove any new annotation that is unsuitable for their model. At any stage, the user may click on the "Annotated Model" tab in Saint, which adds all new annotation to the original model and presents the new SBML model for viewing and download.
While there are a number of tools available for manipulating and validating SBML (e.g. LibSBML), simulating SBML models (e.g. BASIS and the SBML Toolbox ), and analysing simulations (e.g. COPASI,), and running modelling workflows (e.g. Taverna ), Saint is the first to provide basic automatic annotation of SBML models in an easy-to-use GUI. The purpose of Saint is to aid the researcher in the difficult task of information discovery by seamlessly querying multiple databases and providing the results of that query within the SBML model itself. By providing a modelling interface to existing data integration resources and, modellers are able to add valuable information to models quickly and simply.
Saint: automated annotation integration environment code is available from SourceForge or you can try it out here.



